Thermo Fisher Scientific › Electron Microscopy › Electron Microscopes › 3D Visualization, Analysis and EM Software › Use Case Gallery
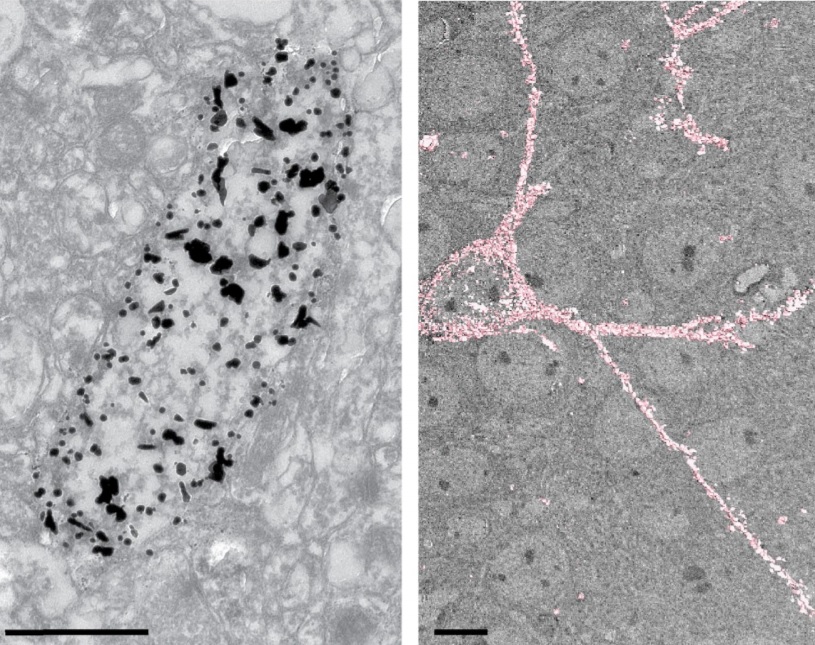
Analysis of neuronal arborization and connections is a powerful tool in fundamental and clinical neuroscience. Changes in neuronal morphology are central to brain development and plasticity and are associated with numerous diseases. Golgi staining is a classical technique based on a deposition of metal precipitate in a random set of neurons. Despite their versatility, Golgi methods have limitations that largely precluded their use in advanced microscopy. We combined Golgi staining with fluorescent labeling and tissue clearing techniques in an Alzheimer’s disease model. We further applied 3D electron microscopy to visualize entire Golgi-stained neurons, while preserving ultrastructural details of stained cells, optimized Golgi staining for use with block-face scanning electron microscopy, and developed an algorithm for semi-automated neuronal tracing of cells displaying complex staining patterns. Our method will find use in fundamental neuroscience and the study of neuronal morphology in disease.
Reconstruction of segmented neurons was done using the Amira software. Segmentation and reconstruction of synapses to impregnated neuron was done by Amira software as well.
For Research Use Only. Not for use in diagnostic procedures.