Thermo Fisher Scientific › Electron Microscopy › Electron Microscopes › 3D Visualization, Analysis and EM Software › Use Case Gallery
In recent years new methodologies and workflow pipelines for acquiring correlated fluorescence microscopy and volume electron microscopy datasets have been extensively described and made accessible to users of different levels. Post-acquisition image processing, and particularly correlation of the optical and electron data in a single integrated three-dimensional framework can be key for extracting valuable information, especially when imaging large sample volumes such as whole cells or tissues.
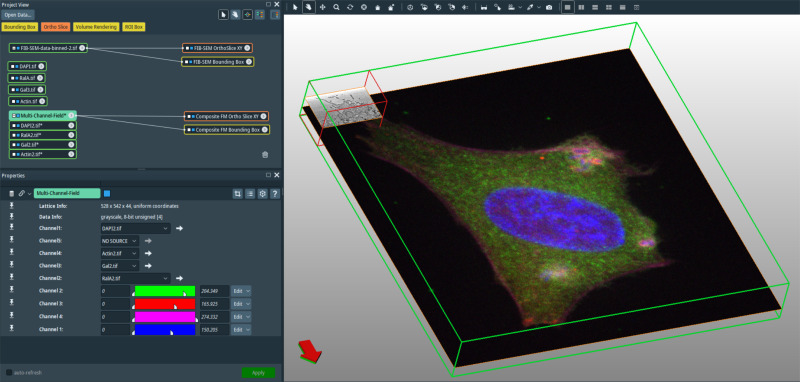
These tasks remain challenging and are often rate-limiting to most users. Here we provide a step-by-step guide to image processing and manual correlation using ImageJ and Amira software of a confocal microscopy stack and a focused ion beam/scanning electron microscopy (FIB/SEM) tomogram acquired using a correlative pipeline. These previously published datasets capture a highly transient invasion event by the bacterium Shigella flexneri infecting an epithelial cell grown in culture, and are made available here in their pre-processed form for readers who wish to gain hands-on experience in image processing and correlation using existing data. In this guide we describe a simple protocol for correlation based on internal sample features clearly visible by both fluorescence and electron microscopy, which is normally sufficient when correlating standard fluorescence microscopy stacks with FIB/SEM data. While the guide describes the treatment of specific datasets, it is applicable to a wide variety of samples and different microscopy approaches that require basic correlation and visualization of two or more datasets in a single integrated framework.
For Research Use Only. Not for use in diagnostic procedures.